Tethered Particle Motion (TPM) uses the Brownian motion
of a polystyrene bead, visible through light microscopy techniques, as a reporter of the motions
of single molecules, which are of course not visible under light microscopes. In our case the
bead is attached via specific small-molecule interactions to one end of a DNA molecule whose other
end is attached via different small-molecule interactions to the surface of a glass coverslip, as
shown below.
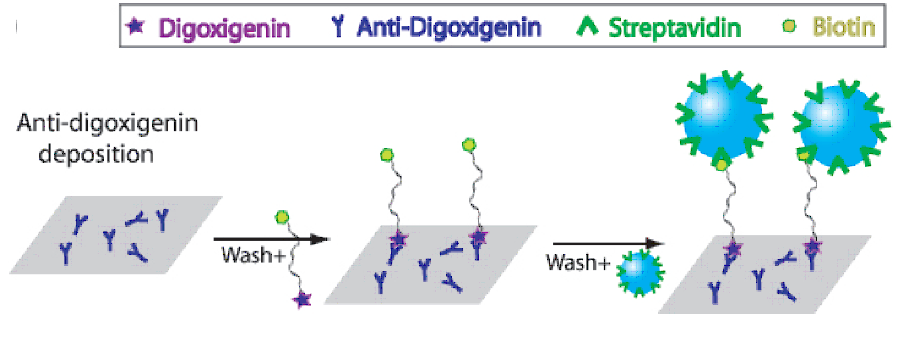
The extent of the bead's motion is related to
the length of the DNA molecule: longer DNA's will allow a greater radius of motion of the attached
bead, while shorter DNA's will make this radius of motion smaller. Thus TPM allows us to study
processes that alter the length of the DNA "tether" by effecting changes in its conformation.
Custom Matlab software allows us to monitor the Brownian motion of the bead as a function of time ("RMS"),
giving a readout in real time of the DNA configuration; we can also histogram the RMS motion to clearly
observe distinct states:

The September '07 Bootcamp TPM project focused on the
formation of loops in the DNA by the Lac Repressor protein, which regulates transcription of the lactose
operon in E. coli. If two or more repressor binding sites (operators) are present on a DNA molecule, the
repressor can bind two operators simultaneously for a certain length of time, forming a loop in the DNA
that effectively shortens the "tether" and therefore reduces the bead's radius of motion. Previous work
in the Phillips lab had characterized the effects of operator spacing, loop sequence, and repressor
concentration on two-operator systems such as the one shown above. However, in E. coli there are actually three operators
present, i.e. the in vivo system has the potential to form three different loops. Biochemical work has
shown that having two sites present increases the efficacy of the repressor's action precisely because
looping can occur; but it has also shown that three operators is even better than two, for unknown reasons (Oehler et al, 1994).
During the September '07 Bootcamp the TPM group examined looping in the three-operator system, with the
in vivo loop spacings and loop sequences, and at a variety of concentrations similar to those
found in vivo, with the exciting result that concentration seems to play an important role in which of
the three loop dominates the system.

The final presentation given on the last day by the students working on this project is shown here:
|
|
|
|

Copyright Phillips Group 2005-2008 |

